Chemical Discovery for Industry, Medical with Carnegie Mellon’s Neural Network Tool
ANI dataset
For industrial and medical companies, a central question in modern chemistry is how to identify and synthesize molecules to create useful applications. Even with high performance computing, the research is expensive both in terms of cost and computer time required for the simulations. The good news is that artificial intelligence (AI) and machine learning (ML) are revolutionizing chemical research and creating tools that can be used in a typical enterprise setting.
The National Science Foundation’s (NSF) Early Science Program (ESP) allows researchers to run simulations on some of the nation’s most powerful supercomputers. In 2019, Olexandr Isayev and his team were awarded an NSF ESP research grant to do chemical research using AI and machine learning on the new Frontera supercomputer at University of Texas Austin’s Texas Advanced Computing Center (TACC), currently the fifth fastest supercomputer in the world and the fastest on a university campus. Isayev is an assistant professor of chemistry at Carnegie Mellon University, and his team’s initial research simulated complex chemical reactions and subsequently extended to molecules in water, as well as reactions on surfaces, and combustion.
“Using AI and ML, our team developed a neural network potential tool named ANI that is trained to solve particular quantum mechanical problems,” Isayev said. “One of the goals in creating ANI was to develop a tool that would be able to predict how a molecule will respond in real-world conditions considering the quantum mechanical behavior when it reacts with other chemical systems.” [editor’s note: ANI is short for Accurate NeurAl network engine for molecular energies, the first general-purpose deep neural network inter-atomic potential.]
“All of our quantum mechanical calculations were enabled by Frontera’s Intel Xeon Scalable processors, which allowed us to take advantage of the latest instructions, like Intel AVX-512. We also leveraged the Intel Math Kernel Library for linear algebra and compilers,” Isayev continued.
He added that “the ANI tool can be used in a variety of areas such as predicting new or stronger materials, developing new drugs, or predicting interaction between a ligand with a protein. Enterprises will not need a supercomputer to do the research. Using AI, our free ANI tool learns quantum mechanics, meaning an enterprise can run simulations on their workstations or even a laptop to predict chemical properties of molecules.”
One of the team’s projects uses neural network models in research on chemical reactions.
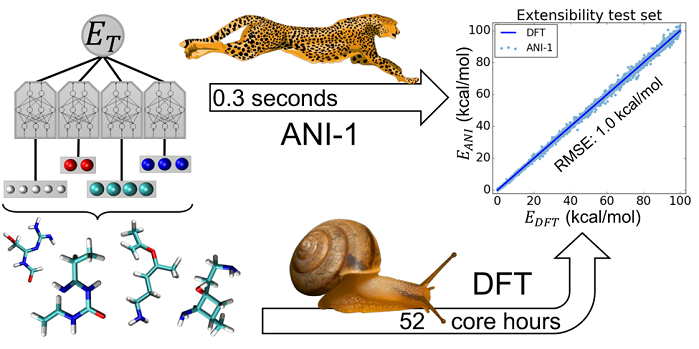
Artistic comparison of ANI neural network and quantum mechanical simulations, courtesy of Olexandr Isayev. https://pubs.rsc.org/en/content/articlelanding/2017/sc/c6sc05720a#!divAbstract
“Neural networks used by our team utilize simple non-linear functions as neurons, and we combine them into a collection of neurons, called layers, that are stacked on top of each other,” said Isayev. “Neural networks loosely mimic how the brain works in processing information. What we are trying to solve is a very complex problem requiring an understanding of the quantum mechanical behavior of many interacting atoms. We are using quantum mechanics (QM) to train the ANI neural network model to understand the molecular behavior of a system and how molecules interact with protein environments. This research can be valuable in providing information to the pharmaceutical industry to design better drugs.” The below image shows an comparison of ANI neural network and quantum mechanics emphasizing speedup while maintain the accuracy.
Carnegie Mellon’s Neural Network Tool
A supercomputer is required to produce the data that trains the neural network ML model. Using Frontera, the researchers generated a large database of molecules with the quantum mechanical energies and properties of each. The research uses quantum mechanical software running on supercomputers to systematically seek the solution to the Schrödinger equation used to predict fundamental properties of molecules. This is a partial differential equation that describes a fundamental quantum-mechanical behavior of atomic or molecular systems and is considered a gold standard for many chemistry applications in terms of accuracy.
“We use supercomputers to solve the equation numerically,” Isayev said. “Using quantum mechanics, and in particular coupled-cluster CCSD(T) approximation is computationally expensive, and often impractical for systems with more than a dozen atoms.”
The initial simulation of their database took 250,000 computer node hours on Frontera and several months to run.
Researchers often use density functional theory (DFT) equations for organic molecules to determine the properties of a many-electron system. DFT simulations use less computer time but the predictions are not as accurate.
The ANI model uses the best of both solutions — the tool first runs a number of DFT calculations to predict system behavior and runs a subset of coupled cluster calculations. This results in simulations that run much faster with high accuracy. “The present work uses transfer learning to train an ML potential that is accurate, transferable, extensible, and therefore, broadly applicable," said Isayev. "In transfer learning, one begins with a model trained on data from a low-fidelity method and then improves the model on data from a high-fidelity method, often yielding high-accuracy predictions even when data are sparsely available,“ he said, adding that the AI-created neural network models have accelerated the computation by six orders of magnitude, or one million times faster.
Isayev said the team used Facebook’s PyTorch framework to generate the neural network model. The ANI model was created as Open Source software and is available for free. The ANI potential is available on GitHub as a user-friendly Python interface integrated with the Atomic Simulation Environment package.
Supercomputers in the Research
The Carnegie Mellon team performed most of their research in creating the database of molecular data on the Frontera supercomputer, which was formally launched in September 2019. The primary computing system is a Dell EMC C6420 system powered by Intel processors with a peak performance of 23.5 petaflops, interconnected by a Mellanox Infiniband HDR and HDR-100 interconnect. Isayev said that quantum mechanics code ORCA and unique code his team runs on Frontera are specifically designed to run well on Intel central processing units (CPU) architectures as well as with Intel Advanced Vector Extensions, Intel Math Kernel Library and Intel compilers. In addition, the memory and storage on Frontera is optimized to eliminate processing bottlenecks. The team worked with TACC staff on storage workflows to run tens of thousands of calculations on Frontera.
The team also uses the Bridges-AI supercomputer at the Pittsburg Supercomputing Center, which is designed to run AI applications and to train neural networks.
“It is important that we democratize the work in science and make tools available for chemists and enterprises who are not specialists in AI but who can benefit from the technologies,” Isayev said. “We have developed our neural network and extremely fast methods with the goal of making these methods available to help transform science and the scientific problems they solve.”
References
1] Teaching Neural Networks Quantum Chemistry, Texas Advanced Computing Center, September 3, 2019, Aaron Dubrow, https://www.tacc.utexas.edu/-/teaching-neural-networks-quantum-chemistry.
2] Approaching coupled cluster accuracy with a general-purpose neural network potential through transfer learning, nature communications, Justin S. Smith, Benjamin T. Nebgen, Roman Zubatyuk, Nicholas Lubbers, Christian Devereux, Kipton Barros, Sergei Tretiak, Olexandr Isayev, and Adrian E. Roitberg, Published in Nature Communications, https://www.nature.com/articles/s41467-019-10827-4
Linda Barney is the founder and owner of Barney and Associates, a technical/marketing writing, training and web design firm in Beaverton, OR. This article was produced as part of Intel’s editorial program with the goal of highlighting science, research and innovation driven by the HPC and AI communities through advanced technology. The publisher of the content has final editing rights and determines what articles are published.











